 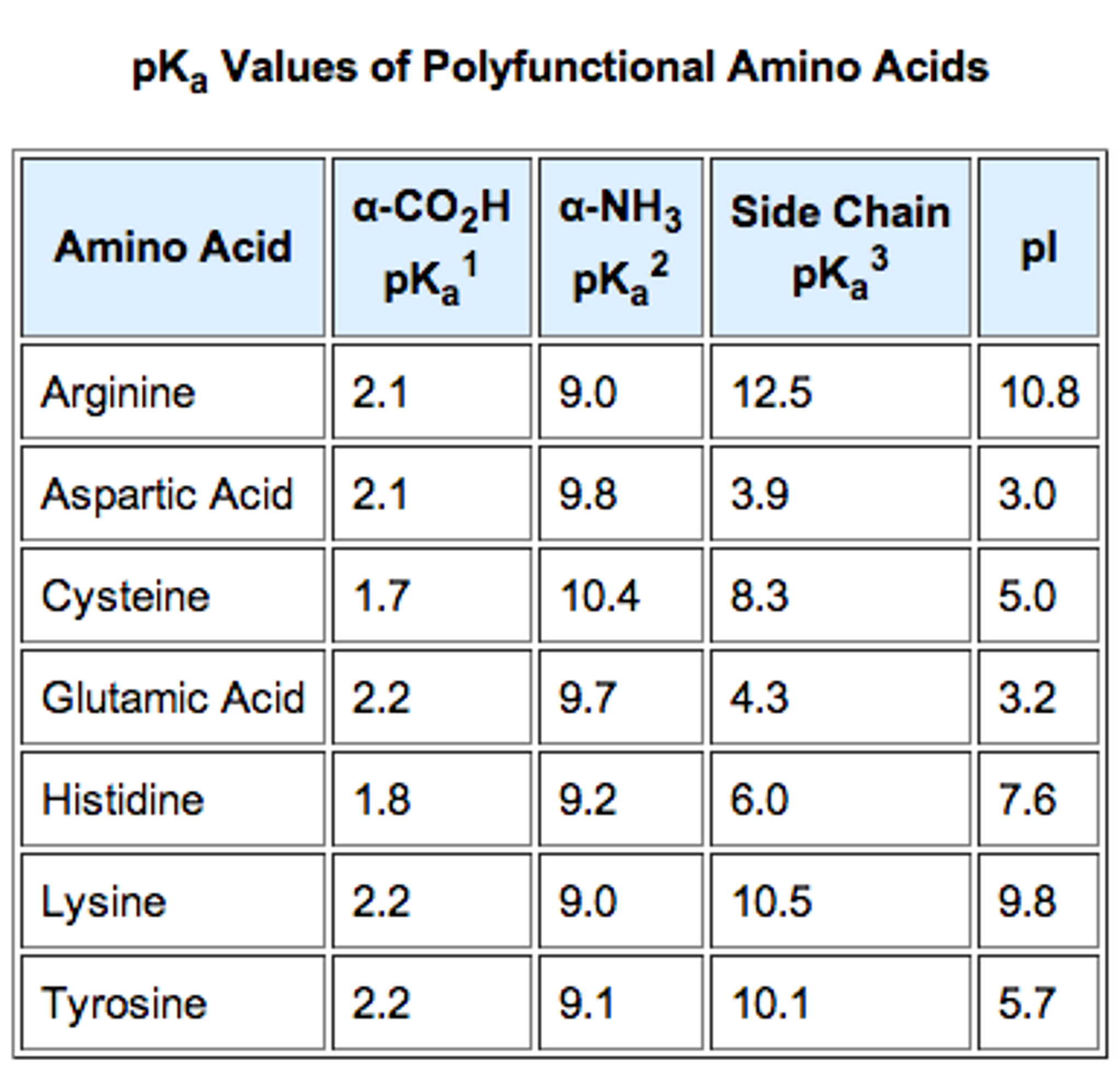
The sequence is ATLDAK and it asks to : Calculate its approximate pI This is what i have so far. approach allow to estimate the isoelectric point of proteins and peptides. I know the general idea behind it but i can't seem to put the nail in the coffin. The latter is determined by the amino acid sequence in the peptide chain. For acidic amino acids, the isoelectric point will be at lower pH as the acidic side chain will introduce an extra negative charge and for basic amino acids, the isoelectric point will be at higher pH as the basic side chain will introduce an extra positive charge. 1 DDS 171 0 How to calculate the pI of a peptide Hello everyone, I am hoping someone can shed some light on this question. A calculator would be used on the first term (to determine the value of 10 -15 ) while, by inspection, it is observed that the last three terms simply reduce to their respective. For neutral amino acids, the side chains are neutral and the isoelectric point is given simply by the average of the pK a values of carboxylic acid and amine. peptide side-chain groups are those commonly reported in. Here are the acid-base equilibria for tyrosine: The form with no net charge is in red (+1 and -1 cancel out to give no net charge). Note: The isoelectric point is given by the average of the pK a values that involve the zwitterions, not just by the pK a values that describe the carboxylic acid group and the amine group. Since the p I is the p H at which the amino acid has no overall net charge, you need to average the p K a values relevant to the protonation/deprotonation of the form with no net charge. Calculate the pI (isoelectric point) of a peptide chain by using the pKa values of the C-terminus, N-terminus, and R groups.DaChemOG ChemMadeSimple pI Is. Since the isoelectric point is given by the average of the pK a values that involve the zwitterion, so we can write the formula for lysine as: In practice you would need to measure it with something like isoelectric focusing, since polypeptides fold and certain groups end up interacting with other groups/being shielded from solvent, meaning you really cant calculate it accurately unless you had structural data and even then it would be extremely difficult. prot pi - protein isoelectric point an online program for calculating pI of proteins (include multiple subunits and posttranslational modifications) CurTiPot a suite of spreadsheets for computing acid-base equilibria (charge versus pH plot of amphoteric molecules e.g. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
